Understanding protein factories: The adventure of deciphering the structure of ribosomes and beyond
This article is based on the discovery of the structure of ribosomes, highlighting the history, and contributions of the scientists who won the 2009 Nobel Prize in Chemistry.

Divyapriya Chandrasekaran
Science Communicator,
Research Matters, Gubbi Labs
04-August-2025
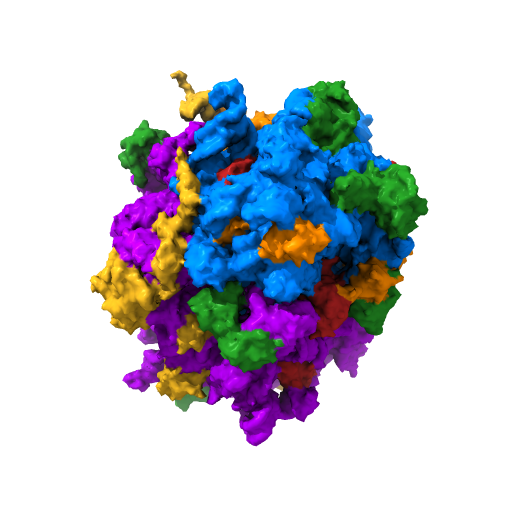
Ribosome of Thermus thermophilus. Credit: https://3d.nih.gov/entries/1331
Cells in our body synthesise a myriad of different proteins within minutes, thanks to small molecular machines called ribosomes—complex molecules composed of ribosomal RNA and proteins. They are found in both prokaryotic and eukaryotic cells. In the eukaryotic cells, they exist in two forms, free in the cytoplasm, and bound to the rough endoplasmic reticulum (RER). They function as factories for protein synthesis.
A ribosome has two subunits-large and small, both joined by magnesium ions. The ribosomes of prokaryotes are composed of 50S (large) and 30S (smaller) subunits, while eukaryotes have 60S and 40S subunits. Today, the ribosome’s atomic structure and function is known, thanks to the discovery by three scientists who shared the Nobel Prize for Chemistry in 2009.
The discovery of molecular machinery
The ribosome was discovered in 1955 by the cell biologist George E. Palade. Using an electron microscope, he identified small globules within a cell. These were initially named ‘Palade particles’ or ‘microsomes’. In 1958, scientists found these microsomes were composed of ribonucleic acid and protein, and renamed them as ‘ribosomes’. They play a crucial role in protein synthesis- in the translation of messenger RNA (mRNA) to proteins. In order to understand their function entirely, it is essential to know their structure and molecular interactions.
Challenges and the discovery of ribosome structure
During protein synthesis, once the ribosomal subunits have assembled, the small subunit of the ribosome binds to mRNA, the blueprint or main ingredient of the recipe. The larger subunit accommodates transfer RNA (tRNA) which adds an amino acid corresponding to the information on the mRNA. Depending upon the length of the mRNA, the amino acid chain extends to form a short or longer protein chain.
Our understanding of the molecular interactions and the three-dimensional structure of ribosomes comes through research using Electron microscopy (EM) and X-ray crystallography. EM magnifies the minute components with greater resolution and is used to study cellular organelles. During the late 1970s, though EM provided overall structure of ribosome, but it was inefficient in rendering precise high resolution images. Hence, scientists had to rely on X-ray crystallography.
In this technique, X-rays hit the crystals of protein molecules, and the scattered X-rays create a diffraction pattern that depicts the arrangement of atoms in the molecule. The mathematical calculations from this help to analyse the pattern for building a protein’s three-dimensional structure.
In spite of this, the isolation and crystallisation of ribosomes remained unmanageable, as earlier painstaking efforts couldn’t render it. The complexity of its structure and distortion due to radiation damage during crystallography had daunted scientists. Hence, understanding the ribosomal structure was considered nearly impossible for decades.
In 1980, Ada E. Yonath, Professor and Director at Weizmann Institute of Science, used cryo-crystallography and exposed the ribosomes under cryo-temperature to successfully obtain its crystals. But, the crystals were of low resolution. After a painstaking 20 years, in 2000, Yonath and her team obtained the high resolution structures of both the ribosomal subunits (30S and 50S) from heat-loving bacteria. Around the same time in 1999-2000, Thomas Steitz, a Professor at Yale University built the accurate structure of the 50S subunit of Haloarcula marismortui—a halophilic archeon from the Dead Sea. Yonath’s team also worked on this archeon in addition to the heat-loving bacterium Thermus thermophilus.
In 1999, Venkatraman Ramakrishnan, a Senior Scientist, from the MRC Laboratory of Molecular Biology, Cambridge, collaborated with other researchers to map the small ribosomal subunit (30S) of T. thermophilus using X-ray crystallography. A year later, his team built the atomic structure of the 30S subunit. In 2007, the team produced the atomic structure of the entire ribosome along with the tRNA and mRNA binding sites.
For their incredible work on mapping the structure of ribosomes, the Nobel Prize for Chemistry 2009, was awarded to Ada E. Yonath, Thomas Steitz, and Venkatraman Ramakrishnan. In his TNQ Distinguished Lectures in 2020, Ramakrishnan shared his journey of solving the mystery of ribosome structure. Knowing the structure of ribosomes was not the end of the story but the beginning of a new era in the research of molecular biology.
The way forward
The structural discovery further aided in understanding the precision of protein synthesis, antibiotic interaction, advances in microscopy, and primitives of ribosomes. This finding led researchers to know how accurately mRNA was read by small ribosomal units, correct tRNA assembly during translation, enabling precise amino acid allocation and peptide bond formation.
The larger subunit binds to the antibiotics, preventing the bacterial protein production and eventually stalling the growth of the pathogen. All three Nobel Laureates had shown antibiotic binding sites in ribosomes. Understanding the antibiotic-binding sites has been helpful in knowing the process of antibiotic resistance and building novel antibiotics to combat it.
